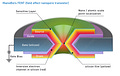
4. INanoBio's FENT: CMOS Transistor Around a Nanopore, for 100X Sensing Speed
My previous post on nanopore platforms as the ultimate future of sequencing has triggered a healthy discussion and debate on Twitter (handle @PreemptDisease). To avoid diverting from the main discussion, I have added a few footnotes. Links to the twitter conversations are provided in the first footnote1. See second footnote2 for additional discussion on commercialization of protein nanopore platforms. I plan to write soon, expanding my thoughts and suggestions on metrics for defining sequencing generations.
Getting back to our discussion - in ion current blockade sensing used by Oxford Nanopore and most others, we effectively count the differential number of ions that slip-past the protein nanopore, as each of the bases (4 or 6-mer strands) transiently occupy the nanopore constriction.
If you think about it, ion current blockade sensing is a surrogate for physically measuring the size of each base relative to the fixed size of the aperture in the protein nanopore, as bases pass through. That is to say, protein nanopore aperture (constriction) can be thought of as a physical ruler to measure and identify the bases, which are sensed and reported by ions.
Ions are the carriers of information in ion-current blockade-based sequencing, whether using protein or solid-state nanopores. As the DNA bases pass through and differentially block the protein pore aperture, ions in the solution transmit the information to the sensing electrodes placed on either side of the nanopore. This generates the sensing ion current, analyzed for base calls. Transmission of information does not occur via physical transport of ions from nanopore all the way to sensing electrodes. It occurs by way of ionic collisions with other ions. The speed of transfer of information via collisions depends on speed of ions (under electrophoretic force and/or electroosmotic flow), rate of collisions and mean free path between collisions, governed by Brownian dynamics. The net speed of ions as information carriers is provided by the characteristic ion mobility in solution.
Ion mobility in aqueous solutions is on the order of 0.001 cm2/ (V·s), which is quite low. Ions are relatively slow or sluggish carriers of information. Due to this, the maximum sequencing speed that can be achieved by ion current blockade sensing on nanopores is around 1000 bases per second, per pore.
We can perform a simple experiment to understand this as detailed in the third footnote3. The cutoff frequency Fc, the operational frequency at which the signal strength falls by 1/2, should be around 10 kHz for ion current sensing (depending on exact platform specs).
The nanopore scientific community simply could not make sense of ion current data to resolve the base information, as DNA was translocating at too fast a speed to be sensed by slow-moving ions in solution. This led to intense research into various methods to slow the DNA transport through nanopore.
Even IBM jumped into the fray with what they called ‘DNA transistor’ to electrostatically slow the DNA speed and read base information, an effort they discontinued after a while. The method that truly succeeded is the polymerase ratcheting approach proposed by Mark Akeson from UC Santa Cruz, which slows DNA from million to <1000 bases per second through the nanopore, as polymerase enzyme processes the DNA strand to make a copy.
One of the biggest advantages of the nanopore approach is the fast DNA translocation through pore, which we lose by slowing it down. This low per-pore sequencing speed is the most significant limitation of Oxford Nanopore and other nanopore sequencing approaches based on ion current blockade sequencing. The other major disadvantages I see are the use of protein pores that increase the complexity of these sequencing platforms, as proteins are inherently unstable, which also makes it challenging to manufacture at scale.
I was following these developments with curiosity, as a part-time postdoc at the time at the Biodesign Institute at the Arizona State University (ASU). Thought, why not turn-the-tables-around on the nanopore sensing problem. Rather than slowing down the DNA speed through nanopore, why not develop a nanopore sensor that’s faster than speed of DNA translocation through nanopore.
The analogy that I often give is: we can see a bullet-train with naked eye but cannot see a bullet fired from the gun, as human eyes are limited to ~ 50 frames captured per second. However, using a DSLR camera with a high frame rate imaging sensor, we can indeed capture a photograph of a bullet traveling in the air. In the same way, if we can make a faster nanopore sensor (instead of slowing down DNA) it becomes possible to sequence DNA at speeds up to million bases per second (and thats what INanoBio is working on).
For this to be possible, we would need a nanopore sensing modality (transducing element) that is based on ultra-fast carriers of information, ideally >100 times faster than speed of DNA through the pore. Since one million bases per second passing through a pore can be thought of as a mega-hertz (MHz) signal, we would need a nanopore sensor capable of 100 MHz sensing speed (Fc > 100 MHz).
My PhD work was on back-gated metal-oxide-semiconductor (MOS) field effect transistor (FET) biosensors. We developed nano-scale FETs and adapted them for the purpose of chemical and biological sensing. The technology is based on standard CMOS semiconductor transistor technology, which forms the basis for all microprocessors and enables operations at over giga-hertz frequencies (billion cycles per second). High speeds are possible because electrons in semiconductor transistors behave as wave particles, and move at speeds that are close to the speed of light. And once fully developed, FET based sensors can be readily produced at established semiconductor manufacturing foundries, that can rapidly scale production while maintaining high fabrication fidelity.
Electrons in semiconductor transistors have mobility on the order of 1000 cm2/ (V·s), which is approximately million times faster than ions in solution. And hence, I reasoned, transistor based nanopore sensors would have orders of magnitude faster speed than DNA passing through a nanopore (at million bases per second), and so, we would not need to slow down the DNA through the nanopore.
We had shown that fully-depleted nano-scale FET sensors are very sensitive to detection of small changes in charge or potential at the sensor surface, reported in peer reviewed publications (ACS Nano and JACS). We demonstrated detection of pyridine molecules in air with ultra-high attomolar sensitivity, and with high detection specificity (ability to discriminate). Pyridine is a small molecule, just a benzene ring with a carbon replaced with nitrogen atom, and smaller than DNA bases. So, we had a sensor platform that can operate at up to giga-hertz speeds, was super sensitive and which can readily be scaled in future - three key features needed to realize ultra-fast nanopore sequencing.
To combine FET transistor sensing technology with a nanopore sequencing approach - I conceived a novel transistor-nanopore sensor termed FENT (field effect nanopore transistor). FENT is a 3D device, a nano-scale vertical transistor surrounding a central nanopore, in 360 degrees. The current in the FENT transistor flows in the silicon channel (inversion current established by biasing buried gate), from top source-contact to bottom drain-contact of the transistor, funneling around the nanopore which acts as a pinch-point.
I got super excited at this novel device idea, and the possibility for ultra-fast sequencing of DNA without slowing it down. I reached out to Dr. Jeff Schloss, the founding director of the Division of Genome Sciences of the Extramural Research Program at the National Human Genome Research Institute (NHGRI). Following his words of encouragement, we submitted an INanoBio-ASU-Stanford proposal to NHGRI’s revolutionary sequencing technology development program, which was awarded. I had given a talk on ultra-fast nanopore sensing paradigms and concepts at the 2015 NHGRI meeting, which incidentally had in the room the founders or CTOs of most of today’s top sequencing companies.
Clicking on the below will lead to conversations on Twitter.

Oxford Nanopore has led the way and shown that applying deep learning and GPU processing on ion-current data makes it possible to achieve 99% base calling accuracy (raw read) using protein nanopores. This was not the case previously, as lack of accuracy was a major barrier for commercialization of nanopore sequencing. Soon, we should expect Roche-Genia, Illumina and others to follow suit with their own protein nanopore sequencing systems, in the next few years. In the case of Illumina, with their NGS systems commanding >75% sequencing market, they are currently under no immediate pressure. However, in my view. this might change in the next 2- 5 year timeframe if Oxford Nanopore and other platforms (including newer NGS platforms) start capturing significant market share. Especially as more Illumina patents expire in the next year or two. It is a question of when and not if.
Place two electrodes in ionic solution, less than millimeter apart. Apply a DC potential (voltage) at electrode 1 and measure ionic current at electrode 2. Now add an AC signal on top of the DC voltage at electrode 1. Measure the current at electrode 2, it will have an AC current signal on top of DC current, which follows the AC voltage signal at electrode 1. Now increase the frequency of AC voltage signal at electrode 1. As frequency increases the amplitude of the AC current signal measured at electrode 2 starts to fall, even while the amplitude of applied AC voltage signal at electrode 1 remains constant. The loss in sensed AC current signal at electrode 1 becomes more dramatic at higher frequencies, as ions in solution fail to respond at high speeds to transmit high-frequency information between electrodes.
The frequency at which the signal strength (power) falls by half, or the 3dB point, is called cut-off frequency (Fc). Students of engineering may be familiar with transfer functions, used to calculate Fc in electronics. Depending on the test setup and experimental parameters, the cut-off frequency for ion current sensing may fall around 10kHz. If we operate above Fc, we dramatically lose sensitivity of ion current sensing of protein nanopore blockade by DNA bases.
Typically, for such sensing applications, we need to sense or sample and acquire data at least ten times (>10x bandwidth) the speed of the events we aim to detect. And that limits ion current blockade sequencing speed to around 1000 bases per second, per pore. So, polymerase ratcheting is used to slow DNA passing through nanopore to 1000 bases per second.